📚 Neural Network Library Implementation
This document reviews the main themes and key takeaways from Deep Learning Systems: Algorithms and Implementation** at Carnegie Mellon University, taught by J. Zico Kolter and Tianqi Chen.
🛠️ I. Introduction & Setup
- 🎉 Welcoming participants back to the Deep Learning Systems: Algorithms and Implementation course.
- 🖥️ Instructions on setting up the Needle repository for lecture 8:
- Cloning the repository.
- Commenting out specific code blocks.
- 📝 Note: Copy-paste the automatic differentiation implementation for this lecture.
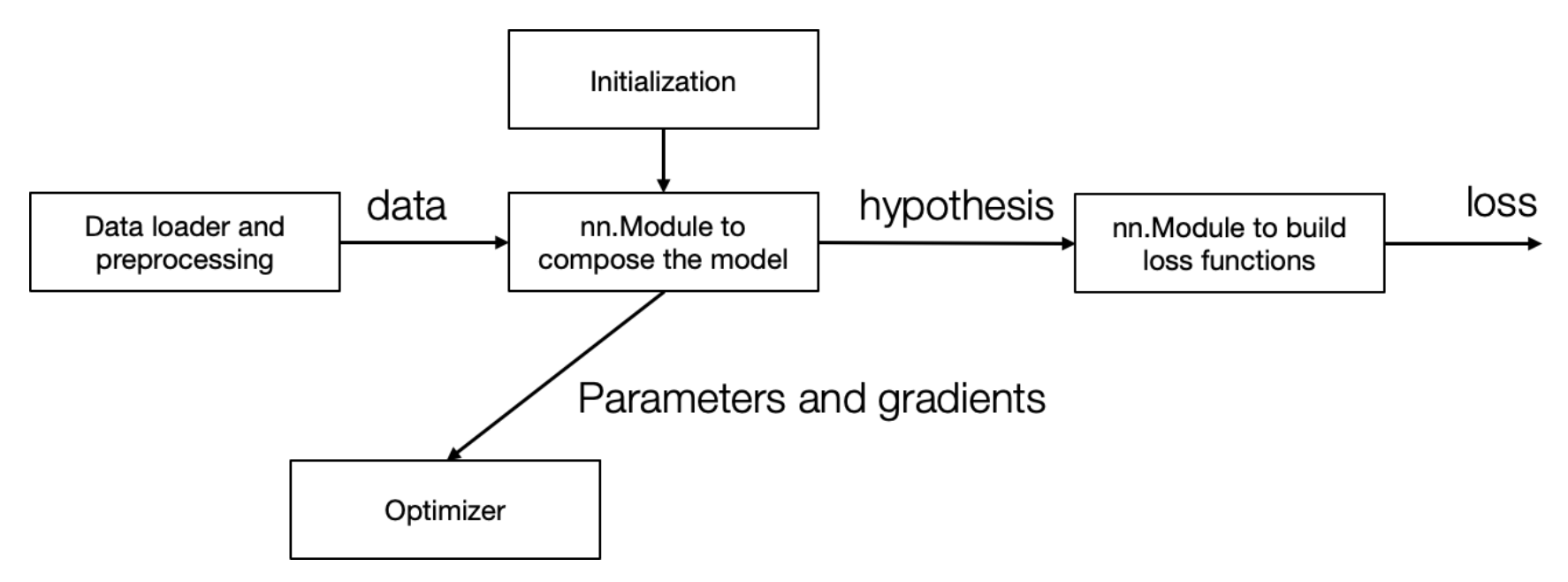
🤔 II. Revisiting Needle & Weight Updates
- 🧠 Refreshing Needle concepts from Lecture 4 and Homework 1:
- Defining Needle tensors and handling weight updates.
- 🚧 Challenges in weight updates:
- Maintaining computational graphs while avoiding memory leaks caused by graph accumulation.
- 💡 Solution: Using detached tensors to eliminate computational graphs during updates:
- Explains
w.dataandw.detachin the Needle codebase.
- Explains
- 🔄 Demonstration:
- Showcasing the benefits of in-place mutations for efficient weight updates.
⚖️ III. Numerical Stability & Softmax
- 🧮 Highlighting numerical instability issues:
- Caused by floating-point precision limits, particularly in softmax calculations.
- 🔥 Example: Overflow errors during naive softmax computation with large values.
- ✅ Solution:
- Subtract the maximum value from the input before softmax to ensure stability.
- 🔍 Similar considerations for:
logsoftmaxandlogsumexp.
🏗️ IV. Building a Neural Network Library: Modules
- 🔨 Transitioning to designing a neural network library:
- Focused on the
nn.Moduleconcept.
- Focused on the
- 🧱 Implementing:
Parameterclass: Distinguishes specific tensors as parameters within modules.Moduleclass: Adds functionality to recursively extract parameters.
- ✨ Demonstration:
- Using the
Moduleclass with aScaleAddmodule:- Defines parameters.
- Implements the forward function.
- Retrieves parameters recursively.
- Modular composition example:
MultiPathScaleAddcombines twoScaleAddmodules.
- Using the
class Parameter(ndl.Tensor):
"""parameter"""
def _get_params(value):
if isinstance(value, Parameter):
return [value]
if isinstance(value, dict):
params = []
for k, v in value.items():
params += _get_params(v)
return params
if isinstance(value, Module):
return value.parameters()
return []
class Module:
def parameters(self):
return _get_params(self.__dict__)
def __call__(self, *args, **kwargs):
return self.forward(*args, **kwargs)
class ScaleAdd(Module):
def __init__(self, init_s=1, init_b=0):
self.s = Parameter([init_s], dtype="float32")
self.b = Parameter([init_b], dtype="float32")
def forward(self, x):
return x * self.s + self.b
class MultiPathScaleAdd(Module):
def __init__(self):
self.path0 = ScaleAdd()
self.path1 = ScaleAdd()
def forward(self, x):
return self.path0(x) + self.path1(x)
Parameter Class
A Parameter is a specialized tensor to represent learnable weights in a model. It extends the base tensor class and is used to distinguish between normal tensors and parameters like weights or biases.
- Purpose: Identifies tensors used as parameters in a model.
- Example: A weight parameter initialized to a specific value is treated as a
Parameter.
_get_params Function
A recursive helper function to extract all Parameter objects from various data structures like dictionaries, modules, or a single parameter.
- Workflow:
- If the input is a
Parameter, it returns a list containing the parameter. - If the input is a dictionary, it recursively extracts parameters from its values.
- If the input is a module, it calls the module’s
parametersmethod to collect all its parameters. - If none of these, it returns an empty list.
- If the input is a
- Example: Extracting parameters from a nested structure or a module simplifies parameter management for training.
Module Class
The base class for all neural network components. It defines methods for parameter retrieval and forward pass invocation.
- Key Methods:
parameters: Recursively collects all parameters from the module’s attributes.__call__: Allows the module to be invoked like a function, automatically calling theforwardmethod.
- Purpose: Provides a unified interface for defining and interacting with neural network components.
ScaleAdd Module
A simple module that implements the operation $( y = x \cdot s + b )$, where $( s )$ (scale) and $( b )$ (bias) are learnable parameters.
- Initialization:
- Takes initial values for $( s )$ and $( b )$ as arguments.
- Registers $( s )$ and $( b )$ as
Parameterobjects.
- Forward Pass:
- Computes the scaled addition operation given an input $( x )$.
- Example:
- Initializes with specific $( s )$ and $( b )$.
- When given an input, it computes the scaled and shifted output.
MultiPathScaleAdd Module
A composite module that combines two ScaleAdd modules, computing the sum of their outputs.
- Initialization:
- Creates two separate
ScaleAddmodules,path0andpath1. - Their parameters are accessible recursively.
- Creates two separate
- Forward Pass:
- Computes the combined output of both
ScaleAddmodules for a given input.
- Computes the combined output of both
- Example:
- Each
ScaleAddmodule applies its own operation to the input. - The outputs are summed to produce the final result.
- Each
Other Module Implementation
Softmax
The Softmax operation applies a numerically stable log-sum-exp function to the input by subtracting off the maximum elements.
$[ \text{LogSumExp}(z) = \log \left(\sum_{i} \exp (z_i - \max{z})\right) + \max{z} ]$
axes: Tuple of axes to sum and take the maximum element over. This uses the same conventions asneedle.ops.Summation().
class LogSumExp(TensorOp):
def __init__(self, axes: Optional[tuple] = None):
self.axes = axes
def compute(self, Z):
### BEGIN YOUR SOLUTION
maxz = array_api.max(Z, axis=self.axes, keepdims=True)
expz = array_api.exp(Z - maxz)
return array_api.log(array_api.sum(expz, axis=self.axes)) + array_api.max(Z, axis=self.axes, keepdims=False)
### END YOUR SOLUTION
def gradient(self, out_grad, node):
### BEGIN YOUR SOLUTION
Z = node.inputs[0]
if self.axes:
shape = [1] * len(Z.shape)
j = 0
for i in range(len(shape)):
if i not in self.axes:
shape[i] = node.shape[j]
j += 1
node_new = node.reshape(shape)
grad_new = out_grad.reshape(shape)
else:
node_new = node
grad_new = out_grad
return grad_new * exp(Z - node_new)
### END YOUR SOLUTION
def logsumexp(a, axes=None):
return LogSumExp(axes=axes)(a)
class SoftmaxLoss(Module):
def forward(self, logits: Tensor, y: Tensor):
### BEGIN YOUR SOLUTION
exp_sum = ops.logsumexp(logits, axes=(1, )).sum()
z_y_sum = (logits * init.one_hot(logits.shape[1], y)).sum()
return (exp_sum - z_y_sum) / logits.shape[0]
### END YOUR SOLUTION
Batch Normalization
$[ y = w \circ \frac{z_i - \textbf{E}[x]}{\sqrt{\textbf{Var}[x] + \epsilon}} + b ]$
Here, the mean and variance refer to the mean and variance over the batch dimensions. The function also computes a running average of mean ($(\hat{\mu})$) and variance ($(\hat{\sigma}^2)$) for all features at each layer. At test time, it normalizes using these quantities:
$[ y = \frac{x - \hat{\mu}}{\sqrt{\hat{\sigma}^2 + \epsilon}} ]$
Batch normalization uses the running estimates of mean and variance instead of batch statistics at test time (i.e., after model.eval() has been called, when the BatchNorm layer’s training flag is set to false).
To compute the running estimates, the following equation is used:
$[ \hat{x}{\text{new}} = (1 - m) \hat{x}{\text{old}} + m x_{\text{observed}} ]$
where $(m)$ is the momentum.
Parameters
dim: Input dimension.eps: A value added to the denominator for numerical stability.momentum: The value used for the running mean and running variance computation.
Variables
weight: The learnable weights of sizedim, initialized to 1.bias: The learnable bias of sizedim, initialized to 0.running_mean: The running mean used at evaluation time, initialized to 0.running_var: The running (unbiased) variance used at evaluation time, initialized to 1.
class BatchNorm1d(Module):
def __init__(self, dim, eps=1e-5, momentum=0.1, device=None, dtype="float32"):
super().__init__()
self.dim = dim
self.eps = eps
self.momentum = momentum
### BEGIN YOUR SOLUTION
self.weight = Parameter(init.ones(self.dim),requires_grad=True)
self.bias = Parameter(init.zeros(self.dim),requires_grad=True)
self.running_mean = init.zeros(self.dim)
self.running_var = init.ones(self.dim)
### END YOUR SOLUTION
def forward(self, x: Tensor) -> Tensor:
### BEGIN YOUR SOLUTION
batch_size = x.shape[0]
feature_size = x.shape[1]
# running estimates
mean = x.sum(axes=(0,)) / batch_size
x_minus_mean = x - mean.broadcast_to(x.shape)
print(mean)
print(mean.broadcast_to(x.shape))
var = (x_minus_mean ** 2).sum(axes=(0, )) / batch_size
if self.training:
self.running_mean = (1 - self.momentum) * self.running_mean + self.momentum * mean.data
self.running_var = (1 - self.momentum) * self.running_var + self.momentum * var.data
x_std = ((var + self.eps) ** 0.5).broadcast_to(x.shape)
normed = x_minus_mean / x_std
return normed * self.weight.broadcast_to(x.shape) + self.bias.broadcast_to(x.shape)
else:
normed = (x - self.running_mean) / (self.running_var + self.eps) ** 0.5
return normed * self.weight.broadcast_to(x.shape) + self.bias.broadcast_to(x.shape)
### END YOUR SOLUTION
Layer Normalization
$[ y = w \circ \frac{x_i - \textbf{E}[x]}{\sqrt{\textbf{Var}[x] + \epsilon}} + b ]$
Here:
- $(\textbf{E}[x])$ denotes the empirical mean of the inputs.
- $(\textbf{Var}[x])$ denotes their empirical variance (using the “biased” estimate, dividing by $(N)$ rather than $(N-1)$).
- $(w)$ and $(b)$ are learnable scalar weights and biases, respectively.
Note: The input to this layer is assumed to be a 2D tensor, with batches in the first dimension and features in the second. The weights and biases may need to be broadcasted before applying them.
Parameters
dim: Number of channels.eps: A value added to the denominator for numerical stability.
Variables
weight: The learnable weights of sizedim, initialized to 1.bias: The learnable bias of sizedim, initialized to 0.
class LayerNorm1d(Module):
def __init__(self, dim, eps=1e-5, device=None, dtype="float32"):
super().__init__()
self.dim = dim
self.eps = eps
### BEGIN YOUR SOLUTION
self.weight = Parameter(init.ones(dim),requires_grad=True)
self.bias = Parameter(init.zeros(dim),requires_grad=True)
### END YOUR SOLUTION
def forward(self, x: Tensor) -> Tensor:
### BEGIN YOUR SOLUTION
batch_size = x.shape[0]
feature_size = x.shape[1]
mean = x.sum(axes=(1, )).reshape((batch_size, 1)) / feature_size
x_minus_mean = x - mean.broadcast_to(x.shape)
x_std = ((x_minus_mean ** 2).sum(axes=(1, )).reshape((batch_size, 1)) / feature_size + self.eps) ** 0.5
normed = x_minus_mean / x_std.broadcast_to(x.shape)
return self.weight.broadcast_to(x.shape) * normed + self.bias.broadcast_to(x.shape)
### END YOUR SOLUTION
🚀 Key Features
- Recursive Parameter Extraction:
- Automatically collects all parameters from nested modules and data structures.
- Simplifies common tasks like weight initialization and optimizer setup.
- Modularity:
- Individual components (like
ScaleAdd) are reusable and composable. - Larger models can be built by combining smaller, well-defined modules.
- Individual components (like
- Forward Pass Handling:
- Abstracted through the
__call__method. - Ensures all modules have a consistent interface for forward computations.
- Abstracted through the
📉 V. Loss Functions & End-to-End Training
- 🎯 Defining loss functions as modules without parameters:
- Example:
L2Loss.
- Example:
- 🔗 Demonstrating end-to-end training:
- Integrates model, loss function, optimizer (SGD), and training loop.
- 🛠️ Highlights:
- Modular approach simplifies component swapping (e.g., hypothesis, optimizer, loss function).
- Contrasts with manual gradient derivation in earlier exercises.
⚙️ VI. Optimizers & Internal States
- 📋 Discussing the Optimizer interface:
- Key functions:
reset_gradient,step.
- Key functions:
- 🚀 Implementing a basic SGD optimizer:
- Structure and update rule.
- 🌀 Advanced optimization methods:
- Adding internal states for features like momentum.
- Briefly introducing momentum variables in update rules.
class Optimizer:
def __init__(self, params):
self.params = params
def reset_grad(self):
for p in self.params:
p.grad = None
def step(self):
raise NotImplemented()
class SGD(Optimizer):
def __init__(self, params, lr):
self.params = params
self.lr = lr
def step(self):
for w in self.params:
w.data = w.data + (-self.lr) * w.grad
x = ndl.Tensor([2], dtype="float32")
y = ndl.Tensor([2], dtype="float32")
model = MultiPathScaleAdd()
l2loss = L2Loss()
opt = SGD(model.parameters(), lr=0.01)
num_epoch = 10
for epoch in range(num_epoch):
opt.reset_grad()
h = model(x)
loss = l2loss(h, y)
training_loss = loss.numpy()
loss.backward()
opt.step()
print(training_loss)
Adam Optimizer
Adam Optimizer
Implements the Adam algorithm, as proposed in Adam: A Method for Stochastic Optimization.
Algorithm
u_{t+1} = β₁ u_t + (1 - β₁) ∇θ f(θ_t)
v_{t+1} = β₂ v_t + (1 - β₂) (∇θ f(θ_t))²
û_{t+1} = u_{t+1} / (1 - β₁^t) (bias correction)
ˆv_{t+1} = v_{t+1} / (1 - β₂^t) (bias correction)
θ_{t+1} = θ_t - α (û_{t+1} / (√(ˆv_{t+1}) + ε))
Important: Ensure bias correction is applied during optimization.
Parameters
params: Iterable of parameters of typeneedle.nn.Parameterto optimize.lr(float): Learning rate.beta1(float): Coefficient for computing the running average of the gradient.beta2(float): Coefficient for computing the running average of the square of the gradient.eps(float): Term added to the denominator for numerical stability.weight_decay(float): Weight decay (L2 penalty).
class Adam(Optimizer):
def __init__(
self,
params,
lr=0.01,
beta1=0.9,
beta2=0.999,
eps=1e-8,
weight_decay=0.0,
):
super().__init__(params)
self.lr = lr
self.beta1 = beta1
self.beta2 = beta2
self.eps = eps
self.weight_decay = weight_decay
self.t = 0
self.m = {}
self.v = {}
def step(self):
### BEGIN YOUR SOLUTION
self.t += 1
for i, param in enumerate(self.params):
if i not in self.m:
self.m[i] = ndl.init.zeros(*param.shape)
self.v[i] = ndl.init.zeros(*param.shape)
grad = ndl.Tensor(param.grad, dtype='float32').data + param.data * self.weight_decay
# m_{t+1}, v{t+1}
self.m[i] = self.beta1 * self.m[i] + (1 - self.beta1) * grad
self.v[i] = self.beta2 * self.v[i] + (1 - self.beta2) * grad**2
# bias correction
m_hat = (self.m[i]) / (1 - self.beta1 ** self.t)
v_hat = (self.v[i]) / (1 - self.beta2 ** self.t)
param.data = param.data - self.lr * m_hat / (v_hat ** 0.5 + self.eps)
### END YOUR SOLUTION
🎲 VII. Initialization & Tuple Values
- 🔧 Weight initialization:
- Setting appropriate standard deviation for Gaussian initialization in linear layers.
- Maintaining variance across layers with ReLU activations.
- 🎁 Bonus: TensorTuple for multiple return values:
- Example: FusedAddScalars operator with tuple output.
- Integration with automatic differentiation.
- 💡 Encouragement:
- Explore extending automatic differentiation to data structures like dictionaries.
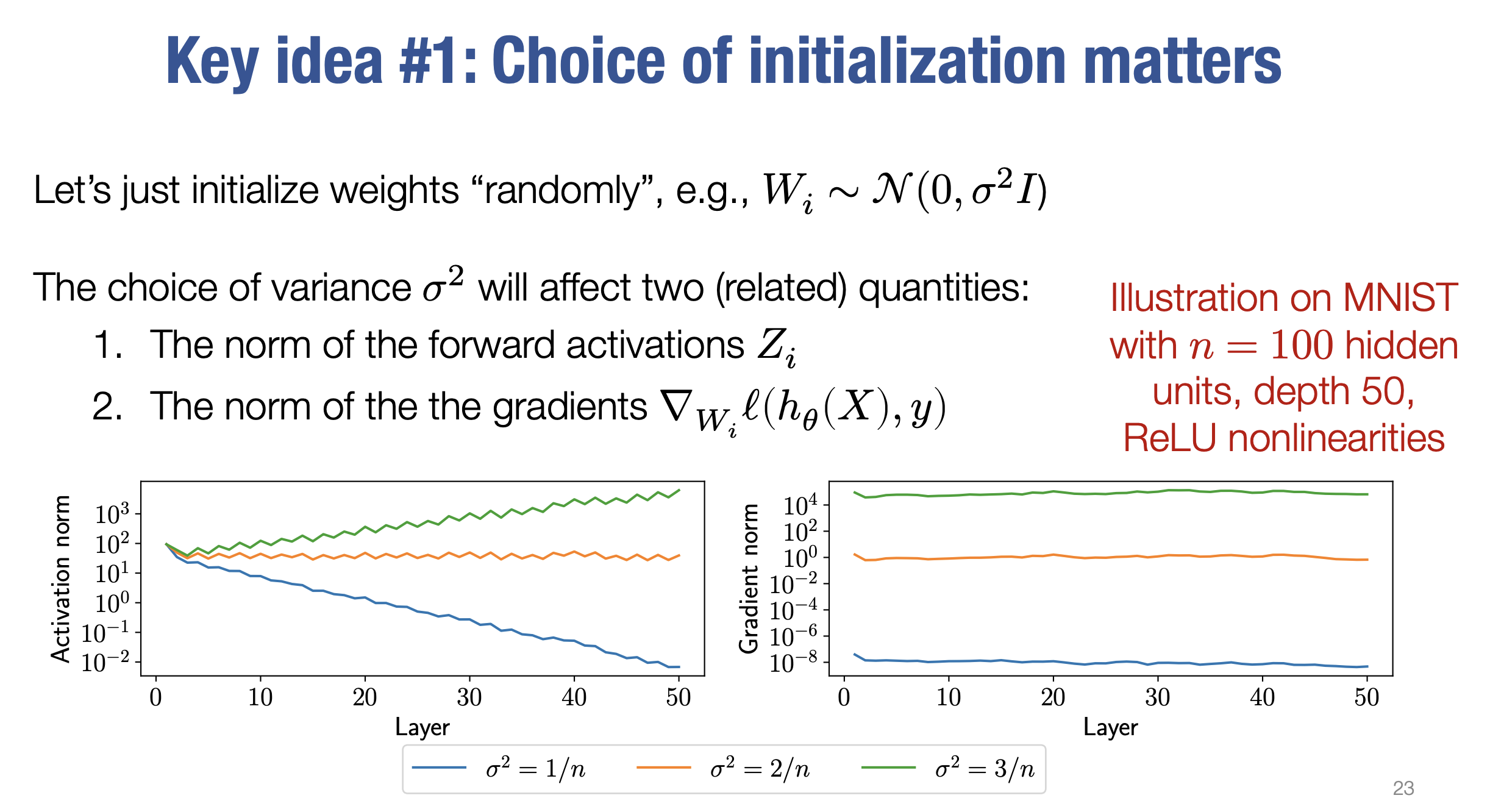
Initialization is crucial for training deep neural networks. Proper weight initialization prevents vanishing or exploding gradients, ensuring stable and efficient training. Two popular methods are Xavier Initialization and Kaiming Initialization.
✨ Xavier Initialization (Glorot Initialization)
📖 Key Idea
Xavier initialization aims to maintain the variance of activations and gradients across layers. It balances the scale of weights to work well with sigmoid and tanh activation functions.
📐 Formula
For a layer with $( n_{\text{in}} )$ input neurons and $( n_{\text{out}} )$ output neurons:
Uniform Distribution
$[ W \sim \mathcal{U}\left(-\sqrt{\frac{6}{n_{\text{in}} + n_{\text{out}}}}, \sqrt{\frac{6}{n_{\text{in}} + n_{\text{out}}}}\right) ]$
Normal Distribution
$[ W \sim \mathcal{N}\left(0, \sqrt{\frac{2}{n_{\text{in}} + n_{\text{out}}}}\right) ]$
🤔 Why It Works
- Ensures the variance of inputs and outputs across layers remains the same.
- Reduces the likelihood of vanishing/exploding gradients in deep networks.
✨ Kaiming Initialization (He Initialization)
📖 Key Idea
Kaiming initialization is designed for ReLU and similar activation functions, which are asymmetric (e.g., they output 0 for negative inputs). It considers the non-linearity of ReLU to maintain proper signal flow.
📐 Formula
For a layer with $( n_{\text{in}} )$ input neurons:
Uniform Distribution
$[ W \sim \mathcal{U}\left(-\sqrt{\frac{6}{n_{\text{in}}}}, \sqrt{\frac{6}{n_{\text{in}}}}\right) ]$
Normal Distribution
$[ W \sim \mathcal{N}\left(0, \sqrt{\frac{2}{n_{\text{in}}}}\right) ]$
🤔 Why It Works
- Accounts for the ReLU activation, which only passes half of the input signal (positive values).
- Prevents the diminishing variance caused by the zero output from ReLU.
🧪 Examples
- Xavier Uniform:
- Used in fully connected layers with sigmoid/tanh.
- Keeps gradients stable during backpropagation.
- Kaiming Normal:
- Common in CNNs with ReLU activations.
- Mitigates gradient flow issues by considering ReLU’s properties.
🚀 Practical Insights
- Use Xavier for layers with symmetric activations like sigmoid/tanh.
- Use Kaiming for layers with asymmetric activations like ReLU.
- Select normal or uniform distributions based on model requirements or defaults from deep learning libraries.
Xavier Initialization (for Sigmoid or Tanh activations) 🌟
- Reason:
- Xavier initialization assumes that both the forward and backward passes of the network should maintain a similar variance.
- Sigmoid and Tanh squash their inputs into small ranges, which can lead to vanishing gradients if the weights are initialized with an improper scale.
- Xavier’s Variance Formula:
- Variance of weights is scaled as:
$[ \text{Var}(W) = \frac{1}{n_{\text{in}} + n_{\text{out}}} ]$ - This balances input and output variances, making it ideal for symmetric activations like Sigmoid and Tanh that don’t “zero out” values.
- Variance of weights is scaled as:
- Best Use Case:
- Works well with symmetric activation functions that preserve the input’s sign, ensuring stable gradients.
Kaiming Initialization (for ReLU or similar activations) 🔥
- Reason:
- ReLU introduces asymmetry by setting negative values to zero, which reduces the number of active neurons (non-zero outputs) in a layer.
- Kaiming Initialization adjusts the weight scale to ensure the variance of activations remains stable across layers, even with ReLU’s sparsity.
- Kaiming’s Variance Formula:
- Variance of weights is scaled as:
$[ \text{Var}(W) = \frac{2}{n_{\text{in}}} ]$ - This accounts for the fact that ReLU “removes” half of the inputs on average, so the factor of $( 2 )$ compensates for the reduced variance.
- Variance of weights is scaled as:
- Best Use Case:
- Ideal for ReLU-like activations to avoid vanishing gradients and ensure efficient learning.
🚀 Why the Difference?
- Xavier focuses on maintaining variance balance for symmetric activation functions like Sigmoid and Tanh, which do not drastically change the input’s magnitude.
- Kaiming compensates for the sparsity introduced by ReLU and similar functions, ensuring that gradients remain meaningful throughout the network.
🎯 Rule of Thumb:
- Use Xavier Initialization for Sigmoid/Tanh.
- Use Kaiming Initialization for ReLU or Leaky ReLU.
def xavier_uniform(fan_in, fan_out, gain=1.0, **kwargs):
### BEGIN YOUR SOLUTION
a = gain * math.sqrt(6 / (fan_in + fan_out))
return a * (2 * rand(fan_in, fan_out, **kwargs) - 1)
### END YOUR SOLUTION
def xavier_normal(fan_in, fan_out, gain=1.0, **kwargs):
### BEGIN YOUR SOLUTION
std = gain * math.sqrt(2 / (fan_in + fan_out))
return std * randn(fan_in, fan_out, **kwargs)
### END YOUR SOLUTION
def kaiming_uniform(fan_in, fan_out, nonlinearity="relu", **kwargs):
assert nonlinearity == "relu", "Only relu supported currently"
### BEGIN YOUR SOLUTION
bound = math.sqrt(2) * math.sqrt(3 / fan_in)
return bound * (2 * rand(fan_in, fan_out, **kwargs) - 1)
### END YOUR SOLUTION
def kaiming_normal(fan_in, fan_out, nonlinearity="relu", **kwargs):
assert nonlinearity == "relu", "Only relu supported currently"
### BEGIN YOUR SOLUTION
gain = math.sqrt(2)
std = gain / math.sqrt(fan_in)
return std * randn(fan_in, fan_out, **kwargs)
### END YOUR SOLUTION
Steps for Xavier Initialization ✨
1. Compute the range (𝑎):
$[ a = \text{gain} \cdot \sqrt{\frac{6}{\text{fan_in} + \text{fan_out}}} ]$
- This range ensures the variance of weights is balanced across layers.
2. Generate uniform random values:
rand(fan_in, fan_out)generates random numbers uniformly in the range $([0, 1])$.- Subtracting 1 and multiplying by 2 shifts the range to $([-1, 1])$.
3. Scale the random values:
- The uniform values are scaled by $(a)$, resulting in weights in the range $([-a, a])$.
Steps for Xavier Normal Initialization ✨
1. Compute the standard deviation (std):
$[ \text{std} = \text{gain} \cdot \sqrt{\frac{2}{\text{fan_in} + \text{fan_out}}} ]$
- The standard deviation ensures that the weights maintain variance across layers.
2. Generate normal random values:
randn(fan_in, fan_out)generates random values from a normal distribution with mean $(0)$ and variance $(1)$.
3. Scale the random values:
- The normal values are scaled by $(\text{std})$, ensuring the correct variance.
